8.3 Processing data
The data processing functionality of patRoon that was discussed before equally applies to IMS workflows. In addition, some extra functionality exists to inspect and filter the data. These are discussed in the next subsections.
8.3.1 Updating feature group properties
The updateGroups() function to update feature group properties (retention time and m/z) was briefly introduced before. This function can also be used to update the assigned mobilities and CCS values to e.g. improve their accuracy after eliminating unwanted features:
8.3.2 Inspecting and plotting data
The plotMobilograms() method function is used to plot extracted ion mobilograms (EIMs), and works very similar as the plotChroms() function introduced before. The function arguments are mostly equal, and the EIMParams argument is similarly used to configure advanced parameters (see ?getDefEIMParams for details).
Most other plotting functions have an IMS argument that controls the inclusion of IMS features and IMS precursors in the plots. This argument is similar as discussed before for e.g. componentization and feature annotation and supports the following values:
IMS=FALSE: Do not plot any IMS features (only supported in post mobility assignment workflows).IMS=TRUE: Only plot IMS features and ignore any IMS precursors.IMS="both": Plot both IMS features and IMS precursors.IMS="maybe": Plot IMS precursors if present, and IMS features otherwise. This is the default value.
The default is "maybe" since most plotting functions primarily use LC-MS data and therefore produce similar plots for IMS features and their IMS precursors.
The spectrumSimilarityIMS() method function calculates the spectrum similarity between MS (or MS/MS) data of a IMS feature and its IMS precursor. This function is internally used by feature annotation if the minIMSSpecSim argument is set, and uses the spectrumSimilarity() function introduced earlier.
The reporting functionality of patRoon contains various functionality that facilitates processing of IMS data.
Some examples are shown below:
#> Using 'mstoolkit' backend for reading MS data.
#> ================================================================================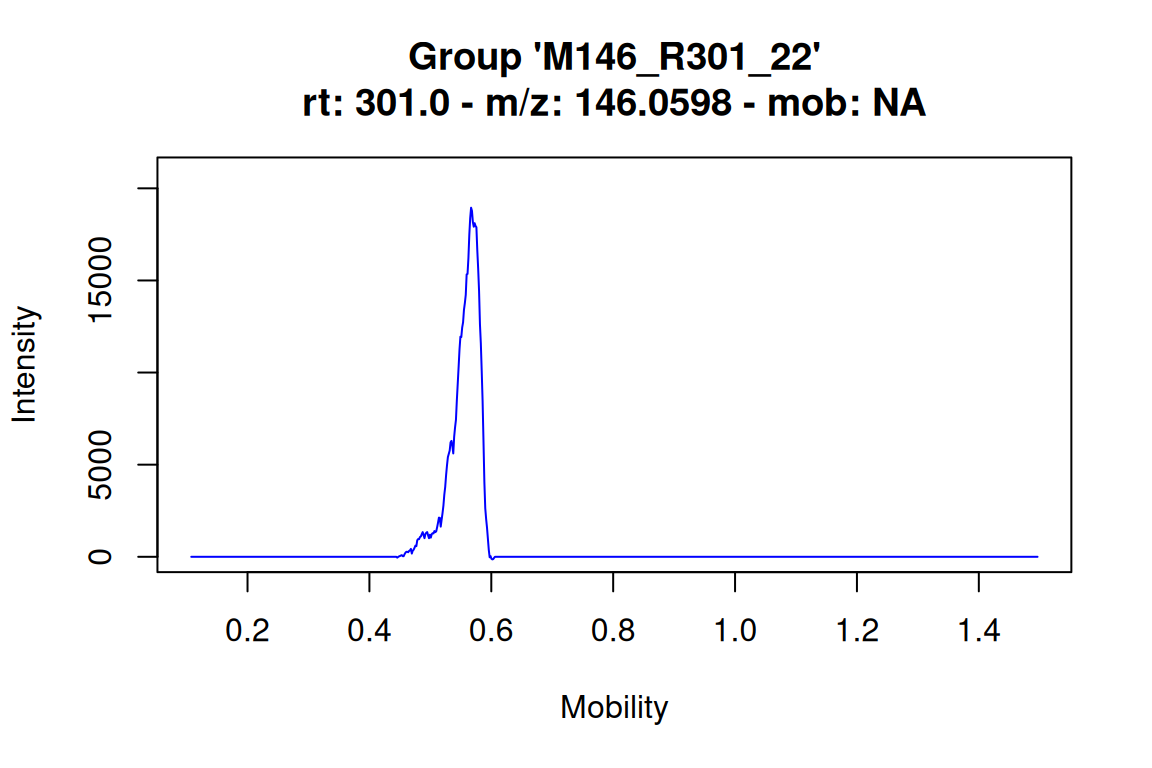
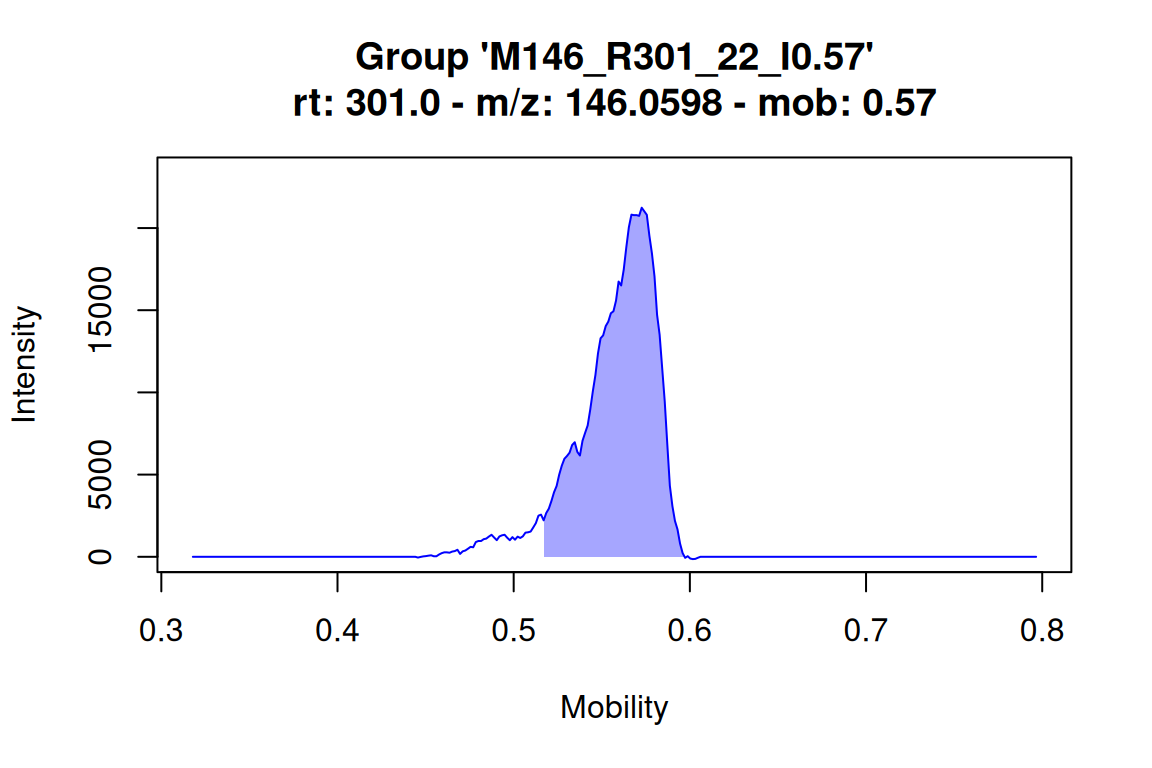
# plot mobilogram for an IMS feature: this highlights the mobility (range) assigned to the feature
# NOTE: the mobility is appended to the feature group name
plotMobilograms(fGroups[1, "M146_R301_22_I0.57"], showPeakArea = TRUE, showFGroupRect = FALSE)#> Using 'mstoolkit' backend for reading MS data.
#> ================================================================================
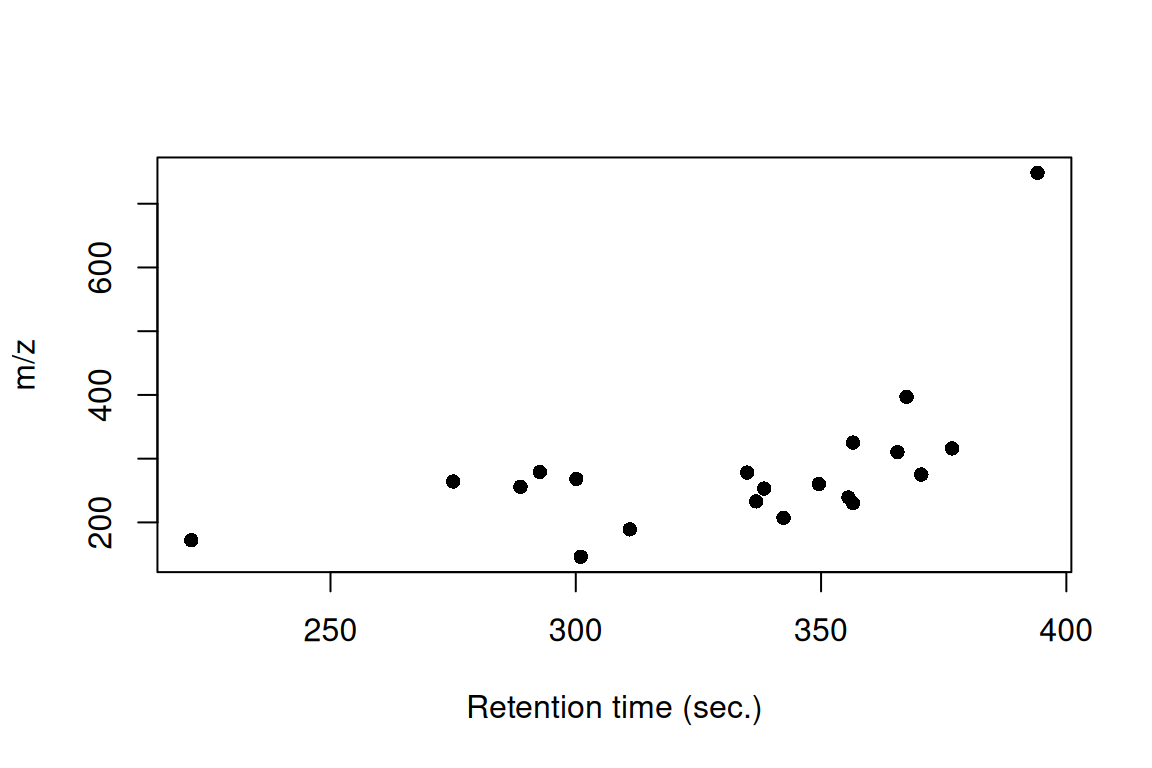
# returns a table with spectral similarities between IMS features and their IMS precursors
spectrumSimilarityIMS(mslists, fGroups)#> group ims_precursor_group similarity
#> <char> <char> <num>
#> 1: M146_R301_22_I0.57 M146_R301_22 0.4591192
#> 2: M172_R222_60_I0.60 M172_R222_60 0.7959959
#> 3: M172_R222_60_I0.62 M172_R222_60 0.8223180
#> 4: M172_R222_60_I0.64 M172_R222_60 0.8559733
#> 5: M189_R311_73_I0.65 M189_R311_73 0.2274158
#> ---
#> 26: M316_R377_214_I0.83 M316_R377_214 0.7105095
#> 27: M316_R377_214_I1.26 M316_R377_214 0.1660779
#> 28: M325_R357_234_I0.87 M325_R357_234 0.5189992
#> 29: M397_R367_277_I0.85 M397_R367_277 0.7907177
#> 30: M748_R394_360_I1.26 M748_R394_360 0.38596878.3.3 Subsetting and filtering data
The following filters are specifically for IMS workflows available:
| Filter | Classes | Remarks |
|---|---|---|
IMS |
featureGroups |
Only keep IMS precursors (IMS=FALSE), IMS features (IMS=TRUE), both (IMS="both") or IMS precursors if available and IMS features otherwise (IMS="maybe"). The IMS argument is also available for the [ operator. |
withIMSPrecursor |
featureGroups |
Only keep IMS features that have an IMS precursor. |
applyIMS |
featureGroups |
Only apply other filters to a subset of features. Should be set like the IMS argument. |
IMSRangeParams |
features, featureGroups, compounds |
Only keep features/annotations within the specified mobility range. The range is configured with the getIMSRangeParams() function. |
IMSMatchParams |
compounds |
Only keep candidates that match reference IMS data. See its description in suspect screening workflows for more details. |
Some examples are shown below:
# keep only IMS features
# NOTE: this effectively results in data that is equal to a direct IMS workflow
fGroupsMob <- filter(fGroups, IMS = TRUE)
fGroupsMob <- fGroups[IMS = TRUE] # same as above
# only keep IMS precursors (post mobility assignment workflows only)
# NOTE: this effectively results in data that is equal to before `assignMobilities()` was called
fGroupsIMSPar <- filter(fGroups, IMS = FALSE)
# perform various filtering steps and ensure that only IMS features with an IMS precursor are kept
fGroupsF <- filter(fGroups, absMinIntensity = 1E5, relMinReplicateAbundance = 1,
withIMSPrecursor = TRUE)
# perform various filtering steps, but not on IMS features
fGroupsF <- filter(fGroups, absMinIntensity = 1E5, relMinReplicateAbundance = 1,
applyIMS = FALSE)
# only keep compound candidates within a defined CCS range
compoundsF <- filter(compounds, IMSRangeParams = getIMSRangeParams("CCS", 150, 200))
# only keep compound candidates with matching CCS (6% tolerance)
compoundsF <- filter(compounds, IMSMatchParams = getIMSMatchParams("CCS", window = 0.06, relative = TRUE))