7.1 Obtaining transformation product data
The generateTPs function is used to obtain TPs for a particular set of parents. Like other workflow generator functions (findFeatures, generateCompounds), several algorithms are available that do the actual work.
| Algorithm | Usage | Input | Output |
|---|---|---|---|
| BioTransformer | generateTPs(algorithm = "biotransformer", ...) |
Parents | TPs structural information |
| CTS | generateTPs(algorithm = "cts", ...) |
Parents | TPs with structural information |
| Library | generateTPs(algorithm = "library", ...) |
Parents (optional), Library (PubChem or custom) | TPs with structural information |
| Formula library | generateTPs(algorithm = "library_formula", ...) |
Parents | TPs with formula information |
| Metabolic logic | generateTPs(algorithm = "logic", ...) |
Feature groups | TPs from m/z differences from pre-defined elemental transformations (based on Schollee et al. (2015)). |
| Formula Annotations | generateTPs(algorithm = "ann_form", ...) |
Parents, formula annotations | Prioritized TPs from annotation candidates (based on Helmus et al. (2025)). |
| Compound Annotations | generateTPs(algorithm = "ann_comp", ...) |
Parents, compound annotaions | Prioritized TPs from annotation candidates (based on Helmus et al. (2025)). |
For most workflows the biotransformer, cts and library algorithms are a good starting point. They are fairly straightforward to use and output TPs with full structural information. The library_formula algorithm may be a suitable alternative if only parents with formula information are available. The ann_comp and ann_form algorithms are meant to elucidate completely unknown TPs, i.e. those that are not predicted or found in literature, and are discussed further below. Finally, the logic algorithm is primarily meant if no or very little information on possible parents and/or TPs is available.
The parent information that is needed for most algorithms is taken from one of the following:
- The data in a suspect list (follows the same format as suspect screening)
- The data from suspects that were matched to feature groups (e.g. obtained with
screenSuspects, see suspect screening) - A
compoundsobject obtained with compound annotation (onlybiotransformer,ctsandlibrary)
The second option is often used in most workflows. The use of compound annotations as parent input could be considered if the parents are not known. However, care must be taken since all the candidates are used, and it is highly recommend to filter the object in advance with e.g. the topMost filter. For library and library_formula, the parent input is optional: if no parents are specified (parents=NULL) then TP data for all parents in the database is used.
An overview of common arguments for TP generation is listed below.
| Argument | Algorithm(s) | Remarks |
|---|---|---|
parents |
biotransformer, cts, library, library_formula, ann_form, ann_comp |
The input parents. |
fGroups |
logic |
The input feature groups to calculate TPs for. |
type |
biotransformer |
The prediction type: "env", "ecbased", "cyp450", "phaseII", "hgut", "superbio", "allHuman". See BioTransformer for more details. |
transLibrary |
cts |
The transformation library that should be used: "hydrolysis", "abiotic_reduction", "photolysis_unranked", "photolysis_ranked", "mammalian_metabolism", "combined_abioticreduction_hydrolysis", "combined_photolysis_abiotic_hydrolysis", "pfas_environmental", "pfas_metabolism". See cts for more details. |
TPLibrary |
library, library_formula |
Custom TP library. |
transformations |
logic |
Custom TP transformation rules. |
generations |
biotransformer, cts, library, library_formula |
The number of transformation generations to consider. |
adduct |
logic |
The assumed adduct of the parents (e.g. "[M+H]+"). Not needed when adduct annotations are available. |
formulas/compounds |
ann_form/ann_comp |
The input formula/compound annotations to extract TPs from. |
TPsRef, fGroupsComps |
ann_comp |
The reference TPs and feature groups to use for candidate ranking. See below for more details. |
TPStructParams |
biotransformer, cts, library, ann_comp |
Other advanced parameters, e.g. to calculate structural similarities. See ?getDefTPStructParams for more details. |
Some examples on how to generate TPs are shown below:
# predict environmental TPs with BioTransformer for all parents in a suspect list
TPsBT <- generateTPs("biotransformer", parents = patRoonData::suspectsPos,
type = "env")
# obtain all TPs from the default library
TPsLib <- generateTPs("library")
# get TPs for the parents matched in a suspect screening
TPsLib <- generateTPs("library", parents = fGroupsScr)
# calculate TPs for all feature groups
TPsLogic <- generateTPs("logic", fGroups, adduct = "[M+H]+")
# use formula annotations to obtain TPs
TPsAnnForm <- generateTPs("ann_form", formulas = formulas)
# use compound annotations to obtain TPs and provide additional suspect and feature data for candidate ranking
TPsAnnComp <- generateTPs("ann_comp", compounds = compounds, TPsRef = TPsBT, fGroupsComps = fGroups)7.1.1 (Custom) Libraries and transformations
By default the library and logic algorithms use data that is installed with patRoon (based on PubChem transformations and Schollee et al. (2015), respectively). However, it is also possible to use custom data. For the library_formula no default library is provided, however, these can easily be generated as is discussed at the end of the section.
To use a custom TP structure library for the library algorithm a simple data.frame is needed with the names, SMILES and optionally log P values for the parents and TPs. The log P values are used for prediction of the retention time direction of a TP compared to its parent, as is discussed later. The following small library has two TPs for benzotriazole and one for DEET:
myTPLib <- data.frame(parent_name = c("1H-Benzotriazole", "1H-Benzotriazole", "DEET"),
parent_SMILES = c("C1=CC2=NNN=C2C=C1", "C1=CC2=NNN=C2C=C1", "CCN(CC)C(=O)C1=CC=CC(=C1)C"),
TP_name = c("1-Methylbenzotriazole", "1-Hydroxybenzotriazole", "N-ethyl-m-toluamide"),
TP_SMILES = c("CN1C2=CC=CC=C2N=N1", "C1=CC=C2C(=C1)N=NN2O", "CCNC(=O)C1=CC=CC(=C1)C"))
myTPLib#> parent_name parent_SMILES TP_name TP_SMILES
#> 1 1H-Benzotriazole C1=CC2=NNN=C2C=C1 1-Methylbenzotriazole CN1C2=CC=CC=C2N=N1
#> 2 1H-Benzotriazole C1=CC2=NNN=C2C=C1 1-Hydroxybenzotriazole C1=CC=C2C(=C1)N=NN2O
#> 3 DEET CCN(CC)C(=O)C1=CC=CC(=C1)C N-ethyl-m-toluamide CCNC(=O)C1=CC=CC(=C1)CTo use this library, simply pass it to the TPLibrary argument:
For library_formula the library follows the same format. However, here the formula should be specified instead of the SMILES with the parent_formula and TP_formula columns (although it is still allowed to only specify SMILES, in which case the formulae are calculated from the SMILES).
For the logic algorithm a table with custom transformation rules can be specified for TP calculations:
myTrans <- data.frame(transformation = c("hydroxylation", "demethylation"),
add = c("O", ""),
sub = c("", "CH2"),
retDir = c(-1, -1))
myTrans#> transformation add sub retDir
#> 1 hydroxylation O -1
#> 2 demethylation CH2 -1The add and sub columns are used to denote the elements that are added or subtracted by the reaction. These are used to calculate mass differences between parents and TPs. The retDir column is used to indicate the retention time direction of the parent compared to the TP: -1 (elutes before parent), 1 (elutes after parent) or 0 (similar or unknown). This is discussed later how this data can be used to filter TP candidates. The custom rules can be used by passing them to the transformations argument:
The genFormulaTPLibrary() utility function can be used to automatically generate TP libraries suitable for the library_formula algorithm. The transformation rules to calculate TPs are specified in the same format as used by the logic algorithm.
myTPFormLib <- genFormulaTPLibrary(parents = patRoonData::suspectsPos, transformations = myTrans)
# also calculate second generation TPs (TPs of TPs)
myTPFormLib2 <- genFormulaTPLibrary(parents = patRoonData::suspectsPos, transformations = myTrans,
generations = 2)
# Use library
TPs <- generateTPs("library_formula", TPLibrary = myTPFormLib)Compared to the logic algorithm, the library_formula algorithm is more (and only) suitable for suspect/target screening workflows, allows multiple transformation generations and allows better customization through manually adding/removing TPs from the library prior to passing it to generateTPs().
7.1.2 TPs from feature annotation candidates
The ann_form and ann_comp algorithms are intended to provide a thorough screening and elucidation of TPs that are otherwise difficult to find with other algorithms. These algorithms assume that TPs of interest can be revealed from a thorough feature annotation workflow based on formulae (ann_form) or compounds (ann_comp). The annotation candidates are prioritized and ranked by the TP Score, which is calculated from properties such as the fit of the candidate structure or formula into the parent (or vice versa) and similarity to suspects (i.e. TPs obtained by other algorithms). For more details see Helmus et al. (2025), ?generateTPsAnnForm and ?generateTPsAnnComp.
To provide a thorough screening with ann_comp, often a large compound database such as PubChem is used for compound annotation. This typically results in tens of thousands of candidates for each parent. Hence, obtaining the compound annotations and generating TPs with ann_comp is computationally intensive and can take multiple hours. Afterwards, the topMost filter should be used to reduce the number of candidates to a manageable size (see the next section).
7.1.3 Processing data
Similar to other workflow data, several generic functions are available to inspect the TP data:
| Generic | Remarks |
|---|---|
length() |
Returns the total number of transformation products |
names() |
Returns the names of the parents |
parents() |
Returns a table with information about the parents |
products() |
Returns a list with for each parent a table with TPs |
as.data.table(), as.data.frame |
Convert all the object information into a data.table/data.frame |
"[[" / "$" operators |
Extract TP information for a specified parent |
Some examples:
# just show a few columns in this example, there are many more!
# note: the double dot syntax (..cols) is necessary since the data is stored as data.tables
cols <- c("name", "formula", "InChIKey")
parents(TPs)[1:5, ..cols]#> name formula InChIKey
#> <char> <char> <char>
#> 1: DEET C12H17NO MMOXZBCLCQITDF-UHFFFAOYSA-N
#> 2: Irgarol C11H19N5S HDHLIWCXDDZUFH-UHFFFAOYSA-N
#> 3: Prometryne C10H19N5S AAEVYOVXGOFMJO-UHFFFAOYSA-N
#> 4: Trimethoprim C14H18N4O3 IEDVJHCEMCRBQM-UHFFFAOYSA-N
#> 5: 1H-benzotriazole C6H5N3 QRUDEWIWKLJBPS-UHFFFAOYSA-N#> name formula InChIKey
#> <char> <char> <char>
#> 1: DEET-TP1 C12H17NO2 FRZJZRVZZNTMAW-UHFFFAOYSA-N
#> 2: DEET-TP2 C12H17NO2 KVTUZBGZTRABBQ-UHFFFAOYSA-N
#> 3: DEET-TP3 C2H4O IKHGUXGNUITLKF-UHFFFAOYSA-N
#> 4: DEET-TP4 C10H13NO FPINATACRXASTP-UHFFFAOYSA-N
#> 5: DEET-TP4 C10H13NO FPINATACRXASTP-UHFFFAOYSA-N
#> 6: DEET-TP5 C4H11N HPNMFZURTQLUMO-UHFFFAOYSA-N
#> 7: DEET-TP6 C8H7O2 GPSDUZXPYCFOSQ-UHFFFAOYSA-M
#> 8: DEET-TP7 C8H9NO WGRPQCFFBRDZFV-UHFFFAOYSA-N
#> 9: DEET-TP1 C12H17NO2 FRZJZRVZZNTMAW-UHFFFAOYSA-N#> name formula InChIKey
#> <char> <char> <char>
#> 1: Irgarol-TP1 C8H15N5S MWWBDLRPMWTLRX-UHFFFAOYSA-N
#> 2: Irgarol-TP2 C11H19N5OS HFCMSBLJLJOGGL-UHFFFAOYSA-N#> parent transformation name_lib
#> <char> <char> <char>
#> 1: DEET Aliphatic hydroxylation of methyl carbon adjacent to aromatic ring / Human Phase I N,N-diethyl-m-hydroxymethylbenzamide
#> 2: DEET Hydroxylation of terminal methyl / Human Phase I N-Ethyl-N-(2-hydroxyethyl)-3-methylbenzamide
#> 3: DEET N-dealkylation of tertiary carboxamide / Human Phase I acetaldehyde
#> 4: DEET N-dealkylation of tertiary carboxamide / Human Phase I N-ethyl-m-toluamide
#> 5: DEET Metabolism Benzamide, N-ethyl-3-methyl-In addition, the following generic functions are available to modify or convert the object data:
| Generic | Remarks |
|---|---|
"[" operator |
Subset this object on given parents |
filter |
Filters this object (available functionality depends on TP generation algorithm) |
convertToSuspects |
Generates a suspect list of all TPs (and optionally parents) that is suitable for screenSuspects |
convertToMFDB |
Generates a MetFrag database for all TPs (and optionally parents, only for TPs with structural information) |
plotGraph |
Generates an interactive plot to explore transformation hierarchies (only for TPs with structural or formula information) |
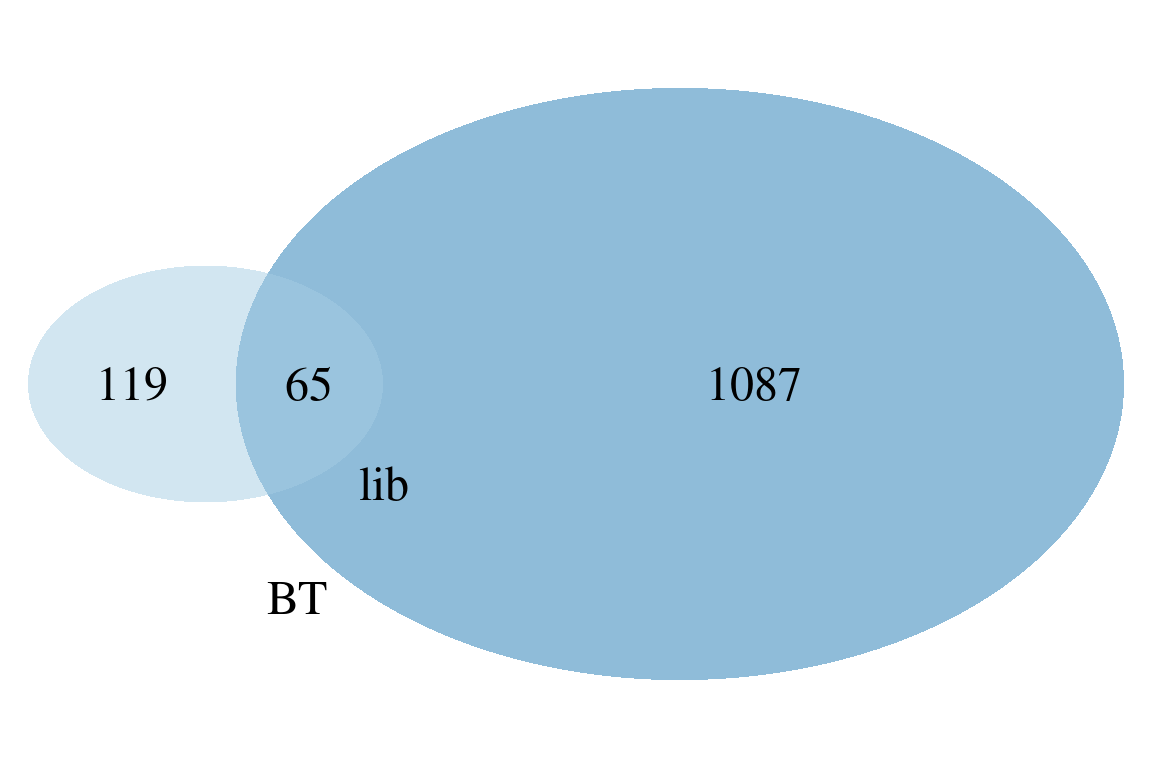
plotVenn, plotUpSet |
Compare results between different algorithms with Venn/UpSet diagrams (only for TPs with structural information) |
consensus |
Combine results from different algorithms (only for TPs with structural information). See the algorithm consensus section. |
The convertToMFDB function is especially handy with predicted TPs, as it allows generating a compound database for TPs that may not be available in commonly used databases. This is further demonstrated in the first example.
TPs2 <- TPs[1:10] # only keep results for first ten parents
# only keep TPs with likely/probably likelihood (specific property for CTS algorithm)
TPsF <- filter(TPs, properties = list(likelihood = c("LIKELY", "PROBABLE")))
# remove transformation products that are isomers to their parent or sibling TPs
# may simplify data as these are often difficult to identify
TPsF <- filter(TPs, removeParentIsomers = TRUE, removeTPIsomers = TRUE)
# remove duplicate transformation products from each parent
# these can occur if different pathways yield the same TPs
TPsF <- filter(TPs, removeDuplicates = TRUE)
# only keep TPs that have a structural similarity to their parent of >= 0.5
# (set TPStructParams=getDefTPStructParams(calcSims=TRUE) when executing generateTPs())
TPsF <- filter(TPs, minSimilarity = 0.5)
# only keep TPs with a TP Score >= 0.5 and the highest 25 TPs per parent
# see ?transformationProductsAnnComp for specific filters
TPsAnnCompF <- filter(TPsAnnComp, minTPScore = 0.5, topMost = 25)
# do a suspect screening for all TPs and their parents
# this is often part of a workflow and is discussed further in the next section.
suspects <- convertToSuspects(TPs, includeParents = TRUE)
fGroupsScr <- screenSuspects(fGroups, suspects, onlyHits = TRUE)
# use the TP data for a specialized MetFrag database
convertToMFDB(TPs, "TP-database.csv", includeParents = FALSE)
compoundsTPs <- generateCompounds(fGroups, mslists, "metfrag", database = "csv",
extraOpts = list(LocalDatabasePath = "TP-database.csv"))